1. Overcoming the challenge of endosomal escape of nucleic acids using DNA-surfactant conjugates
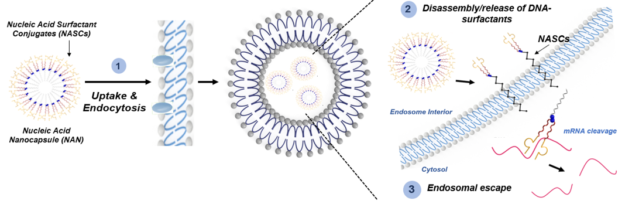
A major part of our research is focused on overcoming the challenge of endosomal entrapment that nucleic acids face during intracellular delivery. Our unique DNA surfactant approach is designed to specifically address this bottleneck in the successful entry of nucleic acids to the cytosol of cells. This area is particularly critical for therapeutic nucleic acids which often require the ability to engage cytosolic targets such as mRNA. Our formulation, which we refer to as a nucleic acid nanocapsule (NAN), is chemically designed to respond to the interior of an endosome. Its degradation and release of nucleic acid cargo into the endosome is tailored to specific enzyme targets and the released oligonucleotide-surfactant conjugate enhances its ability to move through lipid membranes. We are actively studying the fundamental interaction of these oligonucleotide conjugates with lipid membranes both in vitro and in cell culture and are developing new chemical strategies to both track and enhance endosomal escape for improved therapeutic outcomes.
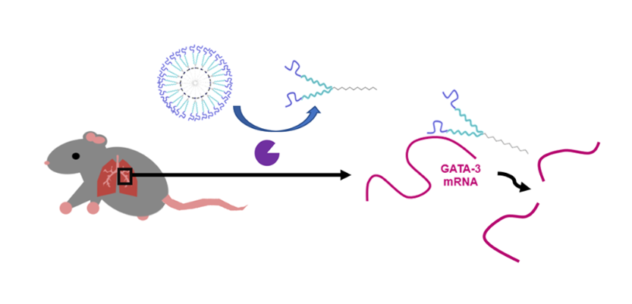
2. Developing biochemically responsive nanomaterials for delivery of therapeutic nucleic acids
New enzyme-responsive nanomaterials are being developed within the group for a variety of applications related to targeted nucleic acid delivery. Current formulations rely on a combination of self-assembled nanomaterials interfaced with enzymatically-assembled ligands. The ligands we utilize include therapeutic oligonucleotides, such as DNAzymes, Ribozymes, siRNA and aptamers. We seek new ways to assemble these biomolecules at organic surfaces that are rapid and scalable. Our goal is to explore the benefits of combining therapeutic oligonucleotides with small molecule drugs to achieve synergistic therapeutic effects in treating diseases. This work is done in collaboration with researchers in the Pathobiology Department at UConn. Current disease models studied include allergically induced asthma.
3. In vitro selection of stimuli-responsive oligonucleotide-nanoparticle conjugates for enhanced multivalent interactions
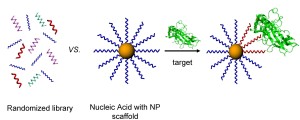
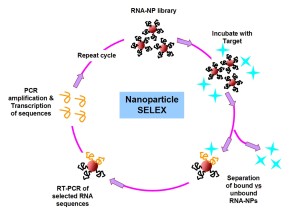
SELEX (systematic evolution of ligands by exponential enrichment) has long been a powerful tool for sorting through enormous libraries of oligonucleotides and identifying stretches of chemical space within them which confer enhanced binding to a given biological target. We are interested in adding a nanoscale scaffold into the selection pressure step as a way to screen for novel multivalent biochemical interactions generated from multi-oligonucleotide contacts. Such contacts are critical for exploring new chemical functionalities of biomaterials at the nanoscale in biologically relevant environments. Successful conjugates will be evaluated for their therapeutic potential in preventing transcription and downstream translation of proteins involved in both inflammation and cancer mechanisms.
In order to assess the possible multivalent interactions that can be made between RNA oriented on a nanoparticle and a given biological target, we will begin by preparing a series of RNA libraries that can specifically probe binding or docking of biomolecules as a function of the RNA-NP conjugate, rather than either component on its own (i.e. unbound RNA or nonspecific interactions with the NP surface). Specifically, we will begin by probing the RNA-NP constructs ability to bind particular transcription factors. Transcription factors are proteins that bind predefined stretches of DNA, while often recruiting other proteins to facilitate transcription and downstream translation of new proteins. Providing a three dimensional architecture for the transcription factors to bind to in the form of the RNA-NP conjugate may be an important method of stalling or preventing downstream mRNA transcription or protein translation, a promising strategy for tackling many disease mechanisms at the genetic level.
Alongside in vitro selection tools, we will take advantage of the large variety of naturally evolved RNAs that have already been identified as having significant therapeutic potential for treating diseases, but which have no clear route to their effective delivery into cells.